Data pre-processing and denoising¶
After the acquisition, we should now have the power doppler signal in a voxels \((z \times x (\times y \text{ if 3D}) \times time)\) , sampled at something like \(0.5\) to \(5Hz\). You can already look at this signal or perform analyses, but some extra steps of preprocessing and denoising might improve signal quality.
Note
It is good practice to measure the effect of each of these steps. If they drastically affect your scientific result is, you should understand why and if keeping these steps, acknowledge the potential confounds. On the contrary, it’s good to keep in mind that artifacts, especially related to the motion of the animal, can affect our results as it is rarely ‘independent’ noise, but often correlated with task variables (in awake animals).
Depending on the type of analysis you want to perform, different types of denoising can be recommended. Here is a list of possible steps to correct for various sources of artifacts. Each step can be carried out independently and none are mandatory. If you are interested in FC analysis check this page addresses more in detail the specific steps you might want to implement.
Motion correction¶
This is to correct for brain deformations that span more than one voxels. Sometimes brain motion can be such that signal from one voxel (eg a visible blood vessel) moves across multiple voxels. In general, the motion is not uniform across the whole brain, rather, it is non-rigid.
-
Example of artefact we will try to correct in this step

Example motion in a awake, head-fixed mouse while the animal moves. Deformations are more visible in deep parts of the brain.
This brain motion is present when animals move and more likely in bigger, stronger animals.
Non-rigid registration can be applied to correct for this kind of motion. There are several toolboxes that implement this process for other imaging modalities (typically 2p imaging) and only need to be adapted/tuned for fUSI.
Tested options include:
Regression of noise¶
Motion can also affect the signal at the sub-voxel level. There might be some residual tissue motion in the power doppler, even though most of it should have been removed by the SVD during pre-processing. To try and mitigate that, we estimate motion-related activity using parts of the images in which we don’t expect any brain-related signal. This can be for example voxels in the gel over the brain, in shadowed parts of the image, or in growth tissue if applicable. From these voxels, we extract the timecourse(s) of artefactual activity (taking the mean, a few PCs, or all voxels). Then, we try to regress this signal out of our signal of interest (i.e. activity in the brain, usually). This can be done with different methods: linear regression, CCA...
Warning
Motion is likely to be correlated to some extent to true hemodynamics signals and/or task variables. Thus, be aware that this step could remove some actual non-artefactual data, especially if your correction procedure is more involved. Depending on your topic of research, you might want or not to remove this movement-related activity, even if biological. In any case, it is good to check whether your correction affect your results and if so, to understand why and make sure it is properly acknowledged.
Outlier removal / Censoring / Scrubbing¶
It is common to have some artefactual, outlier frames. We usually call a frame artefactual if the whole frame (in and out of the brain) has a much different signal than all neighboring frames. Hemodynamics are such that the signal should be somewhat smooth in time, and these frames are violations of this principle. You can find below a few examples of frames to reject
- Flash Artefact: One of the most famous fUSI artefact is the “flash artefact” or “clutter artefact”. It is usually associated with a movement of the animal which induces a change in the spectral (in terms of eigenvalues) distribution of ultrafast ensembles (see PD acquisition) and subsequently a failure of the \(SVD\) based clutter filter. This artefact is characterised by a massive increase of estimated \(PD\) in all voxels (mostly the ones surrounding the brain) as showed here:
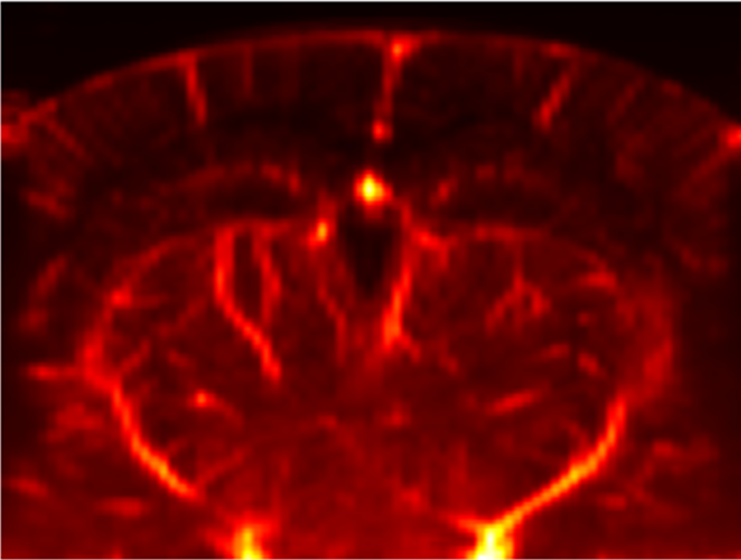
-
Flash artefact video
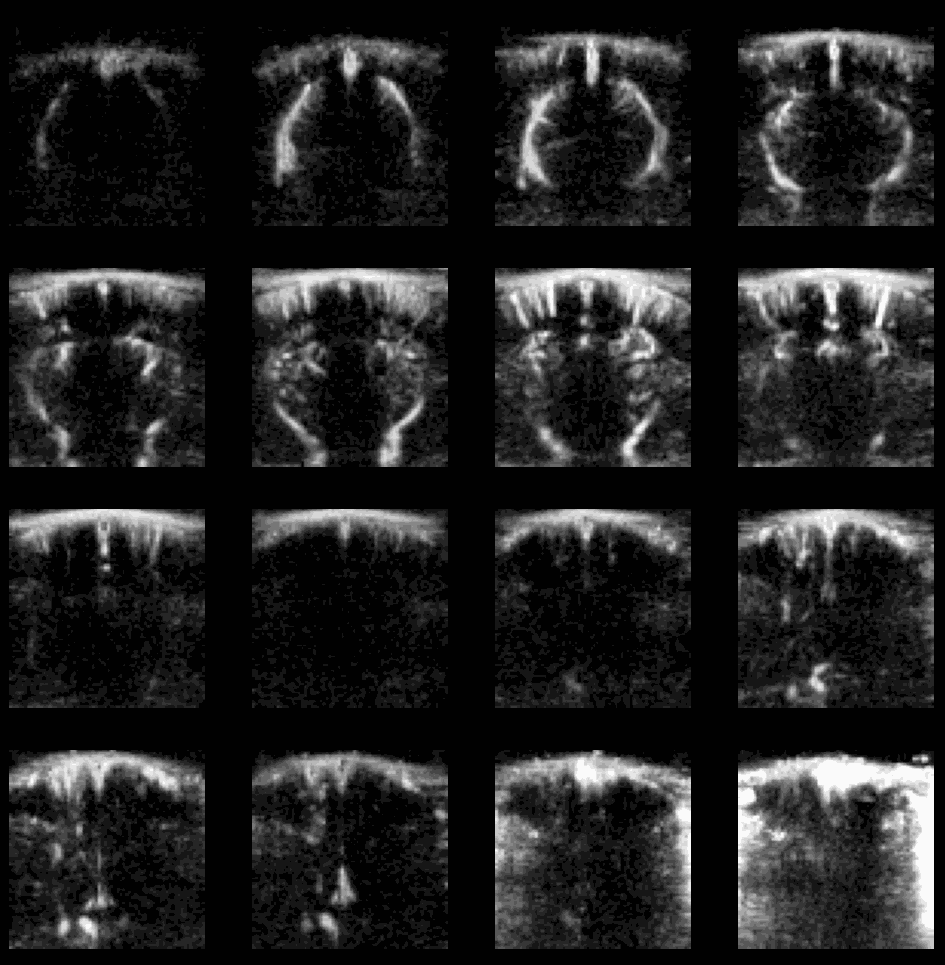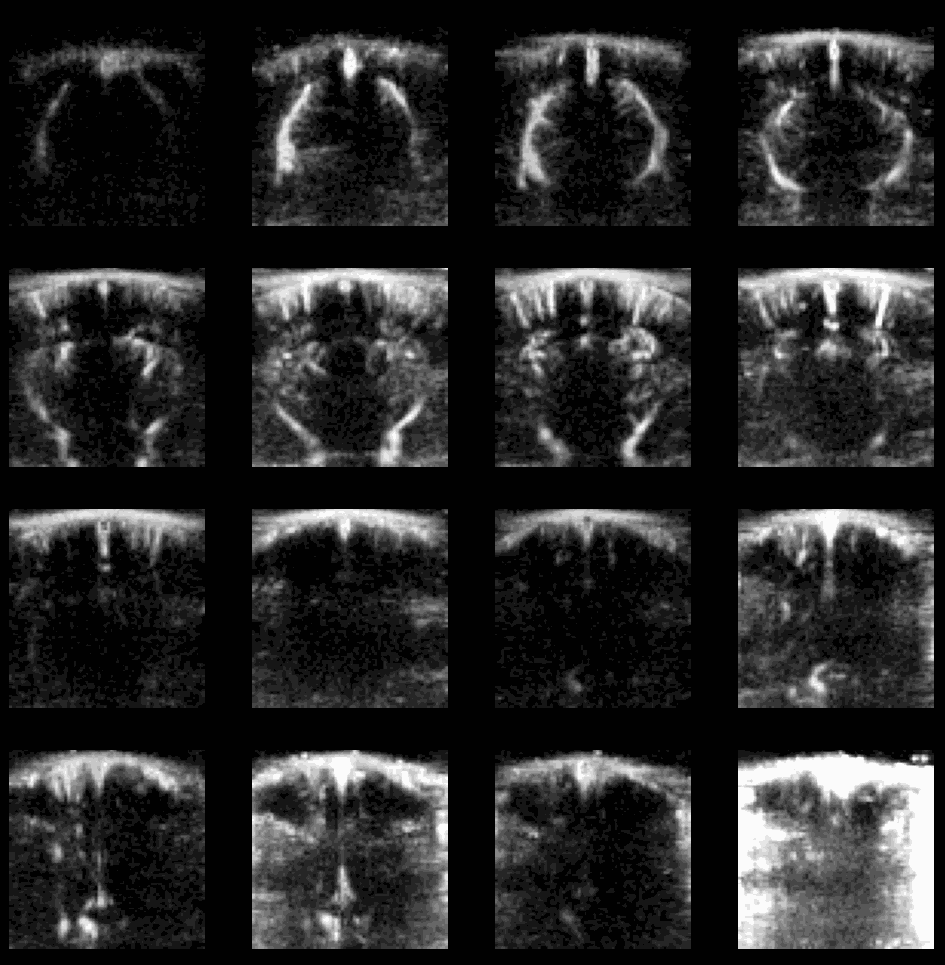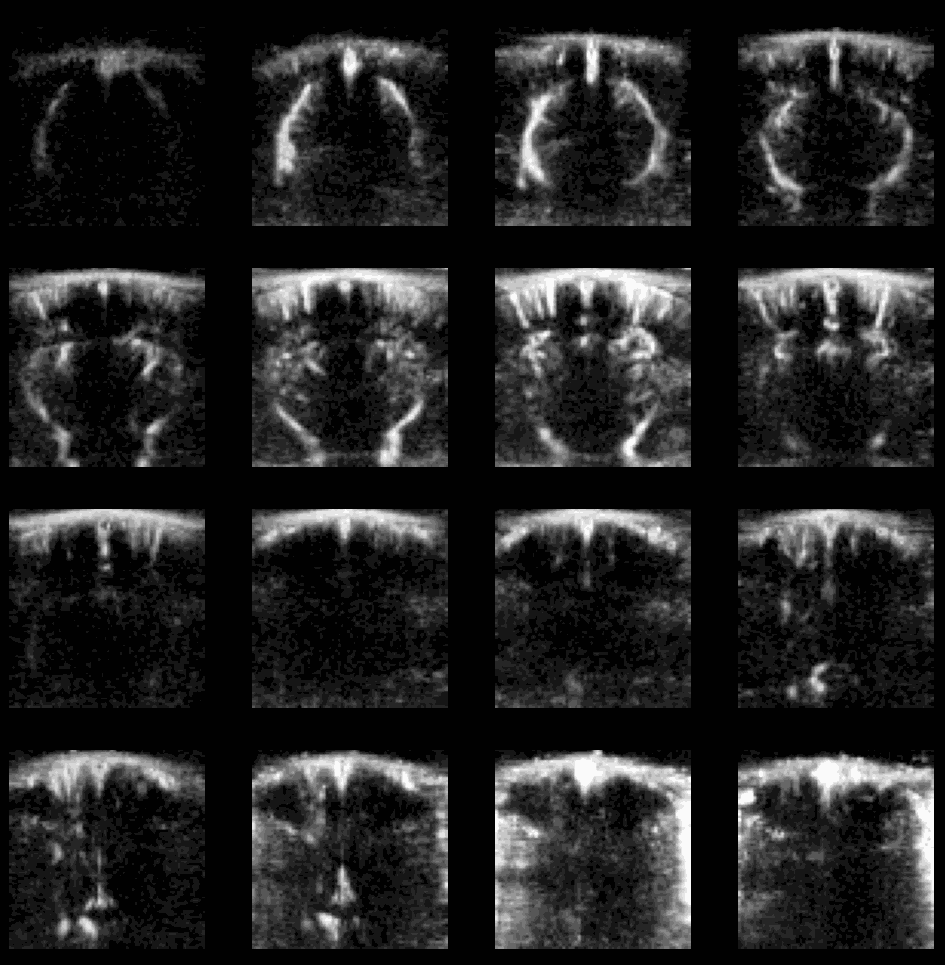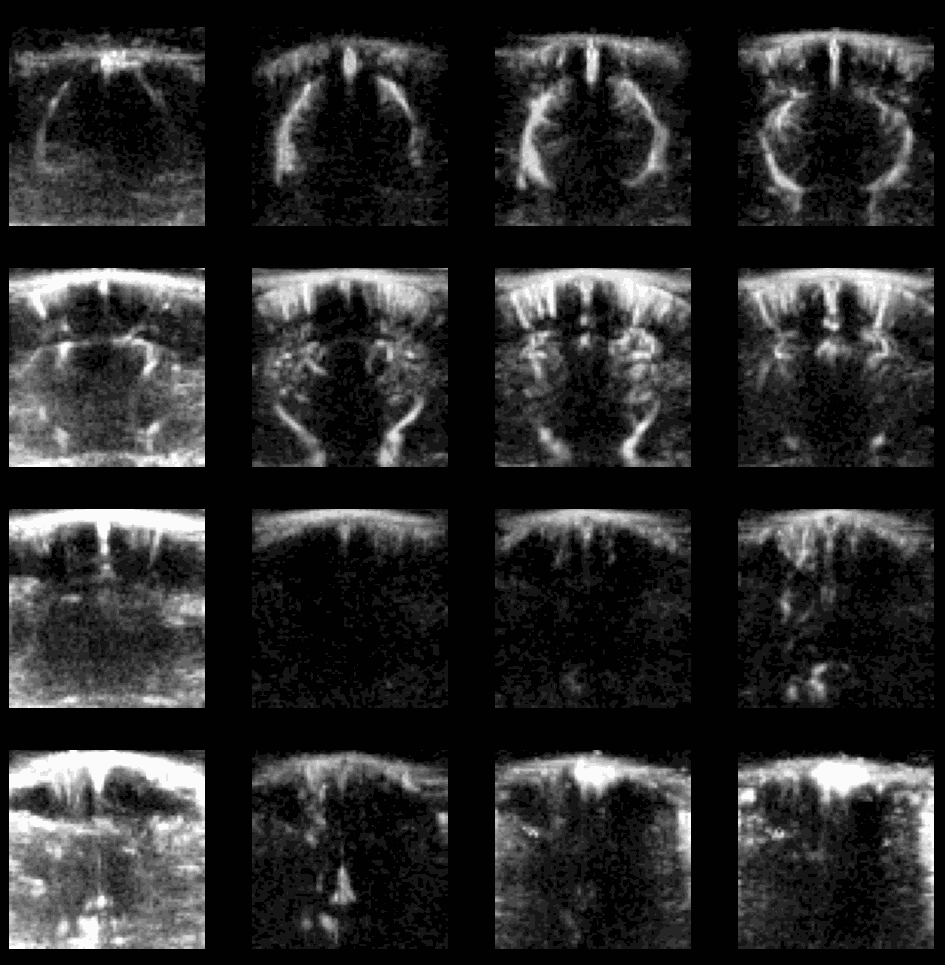
Example of flash artefact in a stacked probe of four linear arrays (lines) acquiring data transcranially at 4 different positions (rows) in an awake mouse, you can see the flashes that affect the all voxels (even in the gel) during movement of the mouse.
To correct for these artefacts, we must first detect them. One way is to simply set a threshold on mean activity (or activity in a subset of voxels, for e.g. in zones with strong noise but little signal), for e.g. 3 standard deviations. Any frame with signal above this threshold will be flagged artefactual and replaced, most simply by a linear interpolation of neighboring frames. Alternatively DVARS (the spatial root mean square of the data after temporal differencing [Smyser et al. 2011], [Power et al. 2012]) has been extensively used to identify local changes in frame intensity associated with such non linear (i.e non rescuable with linear regression) outlier frames. Finally we mention here that Principal Component Analysis (PCA) or Independent Component Analysis (ICA) and their derivates can be used to identify specific timeseries from your whole image, timeseries on which DVARS or simple thresholding can be estimated to better identify images to scrub out.